Efficacy of the modified carbapenem inactivation method (mCIM) for detecting carbapenemase-producing Pseudomonas aeruginosa in a 1,000-bed tertiary hospital in Thailand
Main Article Content
Abstract
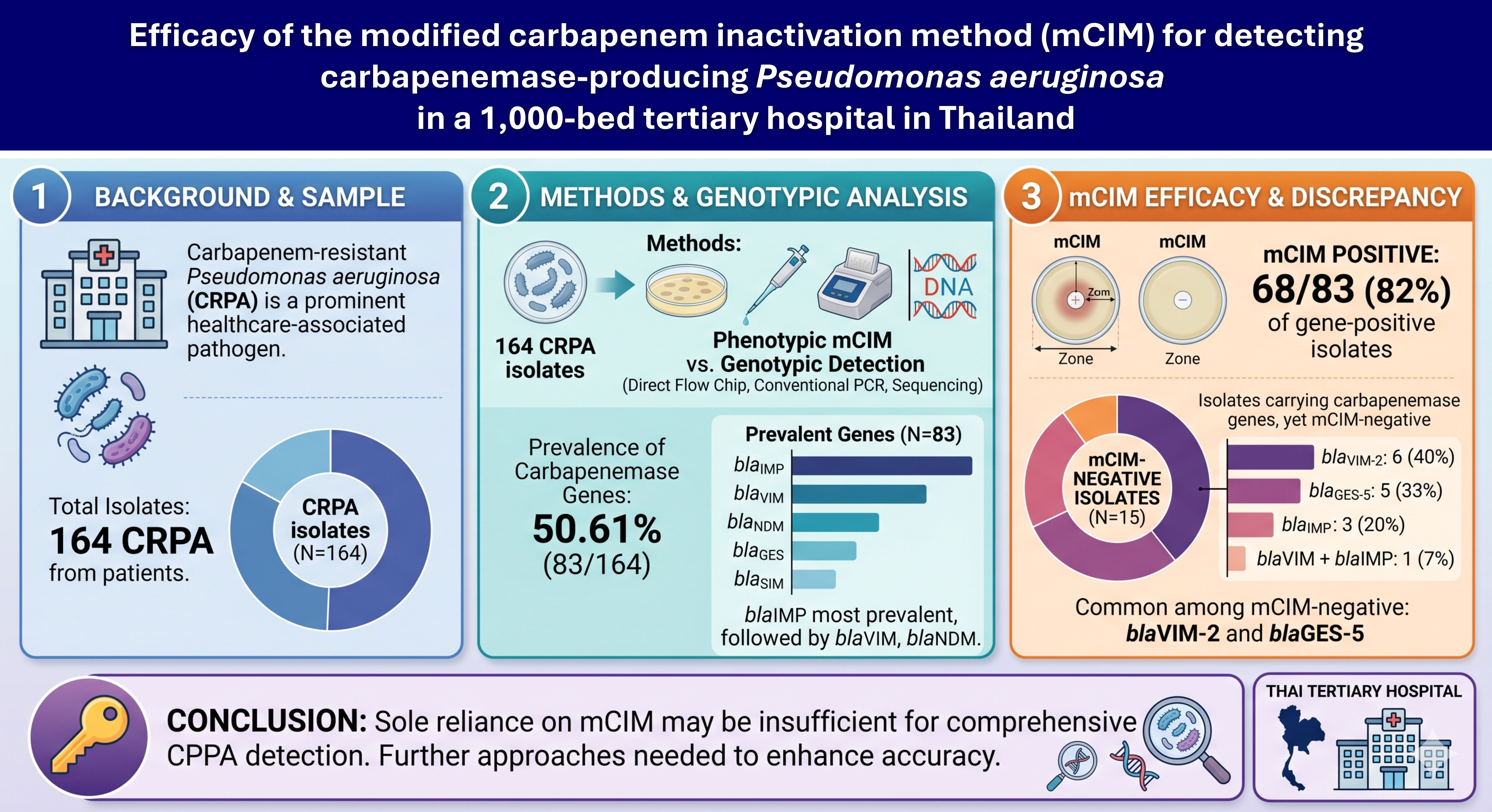
Background: Carbapenem-resistant Pseudomonas aeruginosa (CRPA) is a prominent healthcare-associated pathogen with worldwide implications for public health. Distinguishing between carbapenemase-producing and noncarbapenemase-producing strains over an extended hospitalization period is vital in assessing the risk of treatment failure in infected patients.
Objectives: To study the prevalence of carbapenemase genes and to evaluate the efficacy of the modified carbapenem inactivation method (mCIM) for detecting carbapenemase production among CRPA isolated from Thailand.
Materials and methods: A total of 164 CRPA isolates were phenotypically tested for carbapenemase production by the mCIM comparing with genotypic detection of carbapenemase genes using the direct flow chip kits and conventional PCR. In addition, carbapenemase genes were identified by sequencing.
Results: The carbapenemase genes, including blaIMP, blaVIM, blaNDM, blaGES, and blaSIM, were found in 83 (50.61%) isolates. The blaIMP showed to be the most prevalent gene, followed by blaVIM and blaNDM, respectively. Of these isolates, 68/83 (82%) of carbapenemase gene-positive isolates were positive by the mCIM assay. The 15 mCIM-negative isolates, six (40%) carried blaVIM-2 gene, while five (33%) isolates contained blaGES-5. Moreover, blaIMP was detected in 3 isolates (20%) and one isolate (7%) of a combination between blaVIM and blaIMP genes. Furthermore, we noted that isolates carrying blaVIM-2 were the most common among the mCIM-negative carbapenemase-producing P. aeruginosa (CPPA), followed by blaGES-5.
Conclusion: This study represents a critical report on using mCIM to detect CPPA. It highlights that relying solely on mCIM may not be sufficient for the comprehensive detection of CPPA. Further approaches are needed to enhance the accuracy of detection in this context.
Article Details

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.
Personal views expressed by the contributors in their articles are not necessarily those of the Journal of Associated Medical Sciences, Faculty of Associated Medical Sciences, Chiang Mai University.
References
Tenover FC, Nicolau DP, Gill CM. Carbapenemaseproducing Pseudomonas aeruginosa-an emerging challenge. Emerg Microbes Infect. 2022; 11 (1): 811-4. doi: 10.1080/22221751.2022.2048972.
Yoon E-J, Jeong SH. Mobile carbapenemase genes in Pseudomonas aeruginosa. Front Microbiol. 2021; 12: 614058. doi: 10.3389/fmicb.2021.614058.
Escandón-Vargas K, Reyes S, Gutiérrez S, Villegas MV. The epidemiology of carbapenemases in Latin America and the Caribbean. Exp Rev Antiinfect Ther. 2017; 15 (3): 277-97. doi: 10.1080/14787210.2017.1268918.
Reyes J, Komarow L, Chen L, Ge L, Hanson BM, Cober E, et al. Global epidemiology and clinical outcomes of carbapenem-resistant Pseudomonas aeruginosa and associated carbapenemases (POP): a prospective cohort study. Lancet Microbe. 2023; 4 (3): e159-e70. doi: 10.1016/S2666-5247(22)00307-2.
Choi YJ, Kim YA, Kim J, Jeong SH, Shin JH, Shin KS, et al. Emergence of NDM-1–producing Pseudomonas aeruginosa sequence type 773 clone: shift of carbapenemase molecular epidemiology and spread of 16S rRNA methylase genes in Korea. Ann Lab Med. 2023; 43(2): 196-9. doi: 10.3343/alm.2023.43.2.196.
Butler-Wu SM, Abbott AN. Is this the carbapenemase test we’ve been waiting for? A multicenter evaluation of the modified carbapenem inactivation method. J Clin Microbiol. 2017; 55 (8): 2309-12. doi: 10.1128/ JCM.00321-17.
Laolerd W, Akeda Y, Preeyanon L, Ratthawongjirakul P, Santanirand P. Carbapenemase-producing carbapenem-resistant Enterobacteriaceae from Bangkok, Thailand, and their detection by the Carba NP and modified carbapenem inactivation method tests. Microb Drug Resist. 2018; 24 (7): 1006-11. doi: 10.1089/mdr.2017.0363.
Lisboa LF, Turnbull L, Boyd DA, Mulvey MR, Dingle TC. Evaluation of a modified carbapenem inactivation method for detection of carbapenemases in Pseudomonas aeruginosa. J Clin Microbiol. 2018; 56 (1): e01234-17. doi: 10.1128/JCM.01234-17.
Tsai Y-M, Wang S, Chiu H-C, Kao C-Y, Wen L-L. Combination of modified carbapenem inactivation method (mCIM) and EDTA-CIM (eCIM) for phenotypic detection of carbapenemase-producing Enterobacteriaceae. BMC Microbiol. 2020; 20 (1): 315. doi: 10.1186/s12866-020-01978-7.
Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. 35th Ed. Wayne (PA): Clinical and Laboratory Standards Institute; 2025. CLSI supplement M100.
Poirel L, Walsh TR, Cuvillier V, Nordmann P. Multiplex PCR for detection of acquired carbapenemase genes. Diag Microbiol Infect Dis. 2011; 70 (1): 119-23. doi: 10.1016/j.diagmicrobio.2010.12.002.
Murugan N, Malathi J, Therese KL, Madhavan HN. Application of six multiplex PCR’s among 200 clinical isolates of Pseudomonas aeruginosa for the detection of 20 drug resistance encoding genes. Kaohsiung J Med Sci. 2018; 34 (2): 79-88. doi:10.1016/j.kjms.2017.09.008.
Yoon EJ, Jeong SH. Mobile carbapenemase genes in Pseudomonas aeruginosa. Front Microbiol. 2021; 12: 614058. doi: 10.3389/fmicb.2021.614058.
Gill CM, Lasko MJ, Asempa TE, Nicolau DP. Evaluation of the EDTA-modified carbapenem inactivation method for detecting metallo-β-lactamase-producing Pseudomonas aeruginosa. J Clin Microbiol. 2020; 58 (6): e02015-19. doi: 10.1128/JCM.02015-19.